A Brief Example
Step 1 - Search
Example packages used for this tutorial are: multiqc and tensorflow. Step 1 is about finding multiqc. Step 4 will cover tensorflow.
Go to anaconda.org and search for multiqc in the search bar:

Exam the search result, click on the card with the bioconda tag. This tag is a "channel" name, representing a online folder location where mamba can find and download the correct packages. The most popular channels are conda-forge and bioconda, and remember to avoid using the main and the defaults channel.

Exam the details on the following page, check the version and the home website information.

On the above screenshot, the installation command needs a modification. The conda part in the red circle needs to be changed to mamba. Do not run any conda command on the supercomputers. The syntax for indicating a channel name can either be bioconda::multiqc or -c bioconda multiqc.
Step 2 - Install
Connect to the CiscoVPN. Open a command line interface on Sol by navigating to sol.asu.edu on your browser and selecting the "Sol Shell Access" option from the "System" menu option. Or by SSHing into Sol using the command ssh <asurite>@sol.asu.edu
- CPU-only
- GPU-aware
[rcsparky@login01:~]$ interactive -p htc -c 4 -t 30 -p lightwork
[rcsparky@sc001:~]$ module load mamba/latest
[rcsparky@sc001:~]$ mamba create -n myENV -c conda-forge -c bioconda python=3 multiqc
[rcsparky@sc001:~]$ source activate myENV
(myENV) [rcsparky@sc001:~]$ pip install tensorflow
[rcsparky@login01:~]$ interactive -p htc -c 4 -t 30 -p lightwork
[rcsparky@sc001:~]$ module load mamba/latest
[rcsparky@sc001:~]$ module load cuda-13.x.x-gcc-x.x.x
[rcsparky@sc001:~]$ mamba create -n myENV -c conda-forge -c bioconda python=3 multiqc
[rcsparky@sc001:~]$ source activate myENV
(myENV) [rcsparky@sc001:~]$ pip install tensorflow[and-cuda]
More information can be found here: Managing Python Modules Through the Mamba Environment Manager
Step 3 - Use / Test
Once the myENV environment is ready, multiqc and tensorflow can be used directly or within a python session/script. Below is for using it in the shell, which is also a good testing method for newly built env.
interactive -p htc -c 4 -t 30
module load mamba/latest
source activate myENV
python
>>> import multiqc
Step 4 - What about pip?
As explained in Python Package Installation Method Comparison, the only correct way to use pip on the ASU supercomputers at the moment, is to use it inside an activated mamba env.
There are some packages can only be found on pypi.org but not anaconda.org, then the only option to install them is via pip, inside an activated mamba env. Notably that the current official installation guide for both tensorflow and others prefers pip.

Step 5 - Jupyter Notebook
After multiqc and tensorflow have been installed to myENV, we want to use this mamba env in the Jupyter Notebook session on the Sol web portal. So we need to make a Jupyter kernel from this mamba env. More details are covered in Preparing Python Environments for Jupyter and here are the example steps:
interactive -p htc -c 4 -t 30
module load mamba/latest
mkjupy myENV "myENV_kernel"
Note that you don't need to activate any environment.
To find and use myENV_kernel
- Log in to the Sol web portal
- On the top bar: Interactive Apps > Jupyter > Fill out request form > Connect to Jupyter
- Inside the Jupyter Notebook: Open a Launcher page > Click on
myENV_kernelicon. It usually shows up as the first cube, in front of the public kernels:
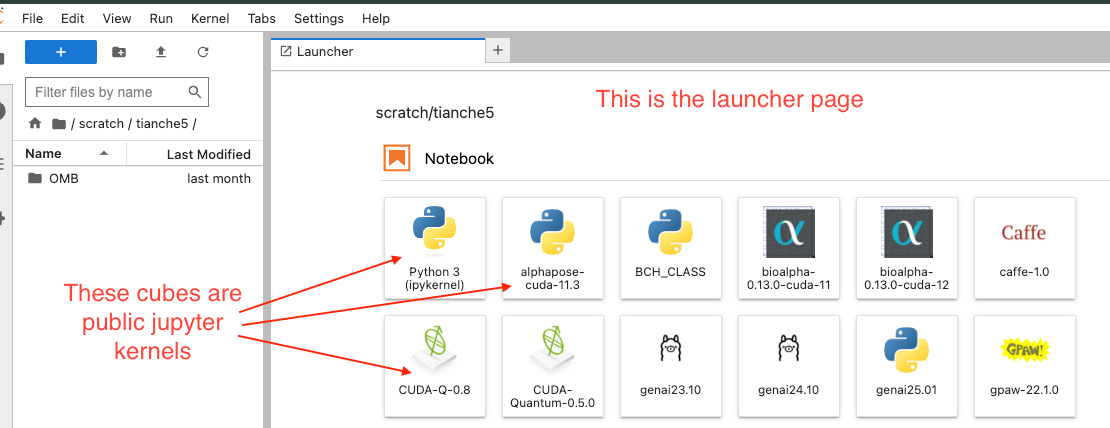
Once a jupyter kernel is made, it cannot be modified. So if you need to add more packages later, the correct steps are:
- Open a shell/terminal to access Sol or Phx
- Add the packages to the existing mamba env
- Recreate the jupyter kernel using the
mkjupycommand. You can use a new name if you want to keep the old kernel. - Launch a new jupyter session on the Web Portal, and look for this new kernel.